pkgs <- c("fs", "futile.logger", "configr", "stringr", "ggpubr", "ggthemes",
"jhtools", "glue", "ggsci", "patchwork", "tidyverse", "dplyr",
"SummarizedExperiment", "jhuanglabRNAseq", "DESeq2", "gplots",
"matrixStats", "Matrix", "dendextend", "ComplexHeatmap", "rstatix",
"clusterProfiler", "DOSE", "org.Hs.eg.db", "EnhancedVolcano",
"GseaVis", "limma")
for (pkg in pkgs){
suppressPackageStartupMessages(library(pkg, character.only = T))
}
setwd("/cluster/home/yjliu_jh/projects/mm/output/response_diff")
# update gene name annotation
hugo_anno <- readr::read_delim("/cluster/home/yjliu_jh/projects/mm/data/hgnc_complete_set.txt",
col_types = cols(intermediate_filament_db = col_character()))
symbol_anno <- hugo_anno %>% separate_rows(prev_symbol, sep = "\\|") |> as.data.frame()
hugo_anno <- hugo_anno[, c("symbol", "ensembl_gene_id", "locus_group")] %>% as.data.frame()
colnames(hugo_anno)[2] <- "gene_id"
# load sampleinfo
merged_sinfo <- read_rds("~/projects/mm/docs/meta/sampleinfo/sampleinfo_jilin_commpass.rds")
resp_info <- merged_sinfo[, c(1:6, 30:33, 111, 143, 285:291)] %>% as.data.frame()
# select samples for comparison
resp_fil <- resp_info[!is.na(resp_info$subtypes), ]
resp_fil <- resp_fil[resp_fil$best_response %notin% "Unknown", ]
# get counts
counts_jilin <- read_csv("/cluster/home/jhuang/projects/mm/analysis/jilin/human/rnaseq/exp/tables/jilin_human_counts.csv")
counts_commpass <- read_csv("/cluster/home/jhuang/projects/mm/analysis/commpass/human/rnaseq/exp/tables/commpass_human_counts.csv")
counts_merged <- cbind(counts_jilin, counts_commpass[, -(1:2)])
counts_merged <- counts_merged[, -2] %>% left_join(hugo_anno) %>% na.omit() %>%
dplyr::select(-c(gene_id, locus_group)) %>% as.data.frame() %>% remove_rownames() %>%
column_to_rownames(var = "symbol") %>% as.matrix() %>% round()
counts_merged <- counts_merged[, resp_fil$sample_id]
#
rna_lib_large <- read_rds("/cluster/home/yjliu_jh/projects/mm/output/rna_lib_mm_samples.rds")
sinfo_comp <- left_join(resp_fil, rna_lib_large)
rownames(sinfo_comp) <- sinfo_comp$sample_id
sinfo_comp <- sinfo_comp[colnames(counts_merged), ]28 Treatment status in this study
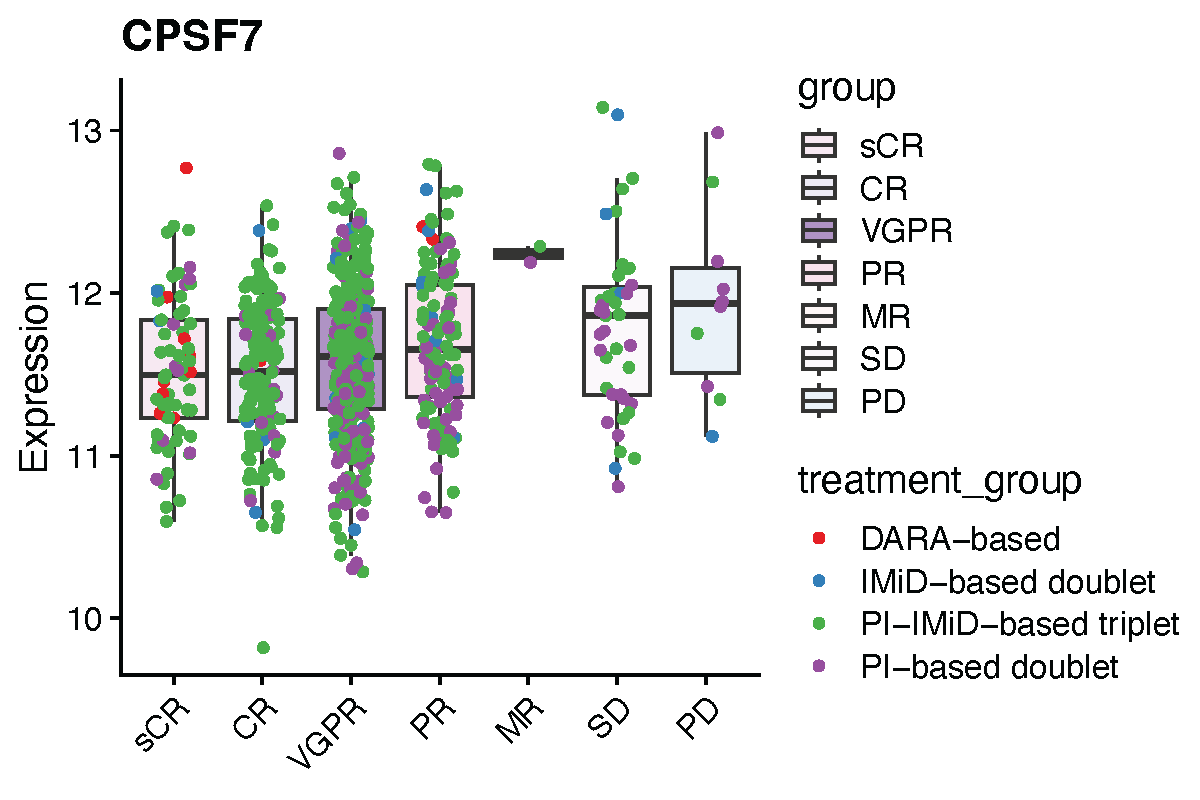
Proteasome inhibitors and immunomodulatory agents are commonly used therapeutic drugs for multiple myeloma (MM). In our dataset, despite the heterogeneity in treatment combinations and regimens, the therapeutic strategies could be broadly categorized into four groups: (1) four-drug regimens based on daratumumab (DARA-based); (2) three-drug regimens combining proteasome inhibitors (PIs) and immunomodulatory drugs (IMiDs); (3) two-drug PI-based regimens; and (4) two-drug IMiD-based regimens. Among these, the three-drug and PI-based regimens were most prevalent, while the IMiD-based regimens were relatively rare. The DARA-based regimens were markedly underrepresented and were therefore excluded from the present analysis.
prepare data: