20 Copy number variation
Copy number variation
20.1 Jilin CNVs
We applied RNAseqCNV to infer arm-level copy number variation (CNV) from bulk RNA-seq samples. For single-cell RNA-seq data, inferCNV was used to estimate CNV profiles at the individual cell level. We observed a degree of concordance between CNV patterns and transcriptomic features, and utilized one of the most prominent CNV-associated expression signatures to define an internal subtype within the HRD group.
20.2 CoMMpass CNVs
In addition, for the CoMMpass dataset, CNV profiles previously reported in published studies were collected and used for comparison. The results demonstrated a high level of concordance in overall CNV trends between our inferences and external data sources.
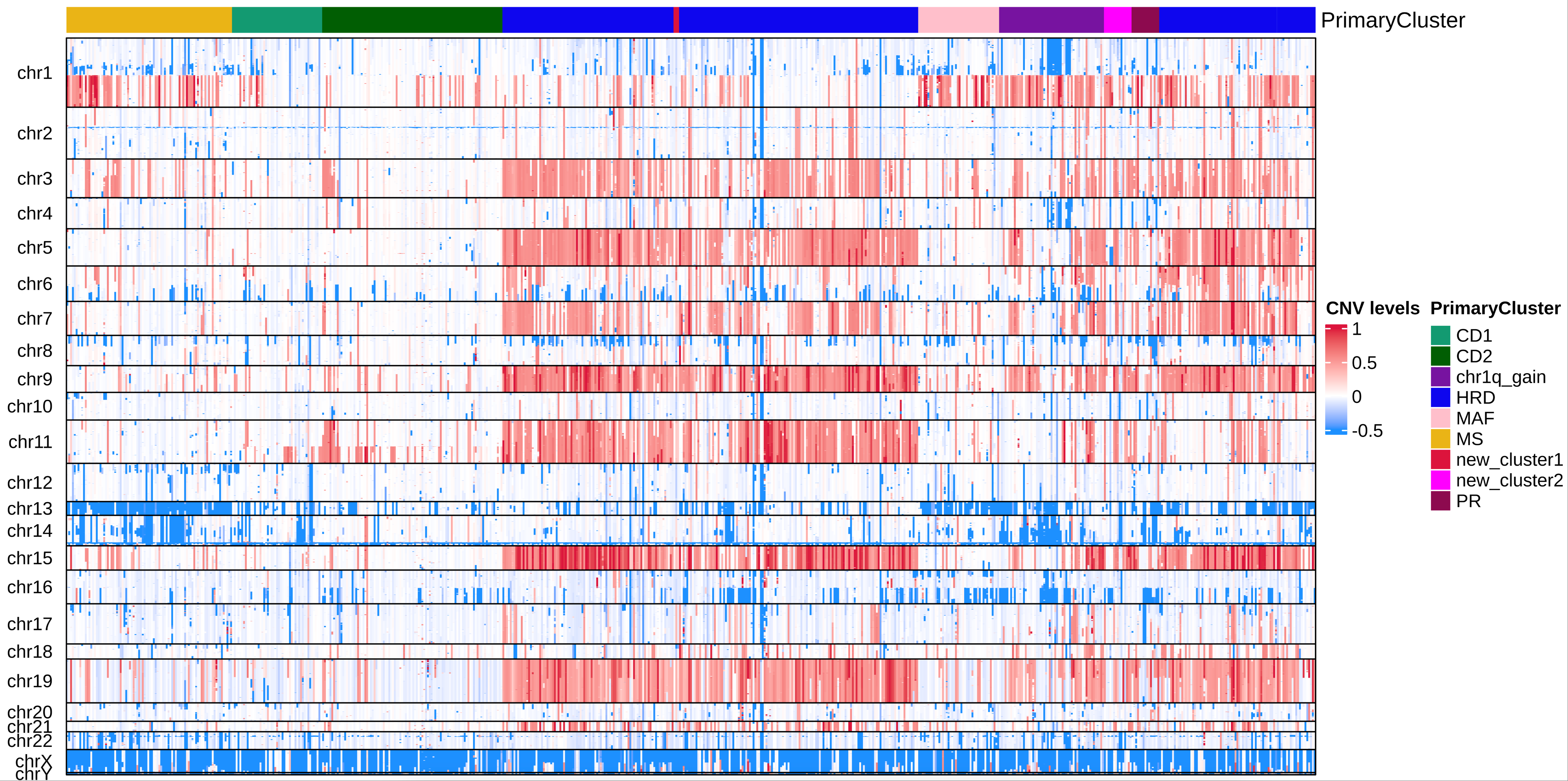
20.3 Total CNV status of MM samples:
A summary of the inferred CNV landscape is shown in the following figure: